

 about
projects
people
publications
resources
resources
visit us
visit us
search
search
about
projects
people
publications
resources
resources
visit us
visit us
search
search
Quick Links
Featured Citations
Bacterial pathogen deploys the iminosugar glycosyrin to manipulate plant glycobiology. Sanguankiattichai N, Chandrasekar B et al. Science. 2025 Apr 18;388(6744):297-303.
Structure of human PINK1 at a mitochondrial TOM-VDAC array. Callegari S, Kirk NS et al. Science. 2025 Apr 18;388(6744):303-310.
Structural pathway for PI3-kinase regulation by VPS15 in autophagy. Cook ASI, Chen M et al. Science. 2025 Apr 11;388(6743):eadl3787.
A coronavirus assembly inhibitor that targets the viral membrane protein. Laporte M, Jochmans D et al. Nature. 2025 Apr 10;640(8058):514–523.
Cytoplasmic ribosomes on mitochondria alter the local membrane environment for protein import. Chang YT, Barad BA et al. J Cell Biol. 2025 Apr 7;224(4):e202407110.
More citations...News
March 19, 2025

|
March 1, 2025
 Follow UCSF ChimeraX on BlueSky!
@chimerax.ucsf.edu
Follow UCSF ChimeraX on BlueSky!
@chimerax.ucsf.edu
December 25, 2024

|
Upcoming Events
UCSF ChimeraX (or simply ChimeraX) is the next-generation molecular visualization program from the Resource for Biocomputing, Visualization, and Informatics (RBVI), following UCSF Chimera. ChimeraX can be downloaded free of charge for academic, government, nonprofit, and personal use. Commercial users, please see ChimeraX commercial licensing.
ChimeraX is developed with support from National Institutes of Health R01-GM129325.
 ChimeraX on Bluesky:
@chimerax.ucsf.edu
ChimeraX on Bluesky:
@chimerax.ucsf.edu
Feature Highlight

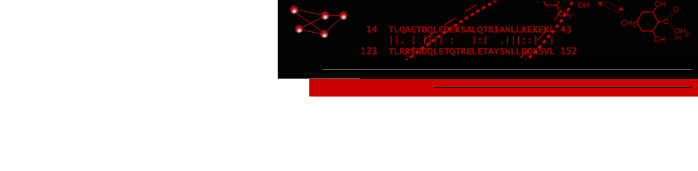
Atomic structures, including cartoons and molecular surfaces, can be colored by the conservation in an associated multiple sequence alignment. The figure shows a structure of the β2-adrenergic receptor signaling complex (PDB 3sn6) with receptor cartoon colored blue→white→red from least conserved to most conserved. The β2-adrenergic receptor is a member of the class A G-protein-coupled receptor superfamily. Conservation was calculated from a superfamily alignment from PASS2 using the entropy-based measure from AL2CO (included with ChimeraX courtesy of Pei and Grishin). The sequence alignment and step-by-step instructions for making this image are given in the Coloring by Sequence Conservation tutorial.
More features...
Example Image

Atomic B-factor values are read from PDB and mmCIF input files and assigned as attributes that can be shown with coloring and used in atom specification. This example shows B-factor variation within a structure of the HIV-1 protease bound to an inhibitor (PDB 4hvp). For complete image setup, including positioning, color key, and label, see the command file bfactor.cxc.
Additional color key examples can be found in tutorials: Coloring by Electrostatic Potential, Coloring by Sequence Conservation
About RBVI | Projects | People | Publications | Resources | Visit Us
Copyright 2018 Regents of the University of California. All rights reserved.